3-Fluoroaniline C6H6FN structure – Flashcards
Flashcard maker : Killian Parsons
Contents
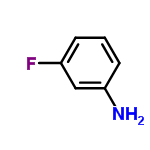
| Molecular Formula | C6H6FN |
| Average mass | 111.117 Da |
| Density | 1.2±0.1 g/cm3 |
| Boiling Point | 186.3±13.0 °C at 760 mmHg |
| Flash Point | 77.2±0.0 °C |
| Molar Refractivity | 30.5±0.3 cm3 |
| Polarizability | 12.1±0.5 10-24cm3 |
| Surface Tension | 39.3±3.0 dyne/cm |
| Molar Volume | 95.9±3.0 cm3 |
- Experimental data
- Predicted – ACD/Labs
- Predicted – EPISuite
- Predicted – ChemAxon
- Predicted – Mcule
- Experimental Physico-chemical Properties
- Miscellaneous
- Gas Chromatography
Predicted data is generated using the ACD/Labs Percepta Platform – PhysChem Module
| Density: | 1.2±0.1 g/cm3 |
| Boiling Point: | 186.3±13.0 °C at 760 mmHg |
| Vapour Pressure: | 0.7±0.4 mmHg at 25°C |
| Enthalpy of Vaporization: | 42.3±3.0 kJ/mol |
| Flash Point: | 77.2±0.0 °C |
| Index of Refraction: | 1.548 |
| Molar Refractivity: | 30.5±0.3 cm3 |
| #H bond acceptors: | 1 |
| #H bond donors: | 2 |
| #Freely Rotating Bonds: | 0 |
| #Rule of 5 Violations: | 0 |
| ACD/LogP: | 1.39 |
| ACD/LogD (pH 5.5): | 1.32 |
| ACD/BCF (pH 5.5): | 5.96 |
| ACD/KOC (pH 5.5): | 124.49 |
| ACD/LogD (pH 7.4): | 1.33 |
| ACD/BCF (pH 7.4): | 6.03 |
| ACD/KOC (pH 7.4): | 125.96 |
| Polar Surface Area: | 26 Å2 |
| Polarizability: | 12.1±0.5 10-24cm3 |
| Surface Tension: | 39.3±3.0 dyne/cm |
| Molar Volume: | 95.9±3.0 cm3 |
Predicted data is generated using the US Environmental Protection Agency’s EPISuite™
Log Octanol-Water Partition Coef (SRC): Log Kow (KOWWIN v1.67 estimate) = 1.28 Log Kow (Exper. database match) = 1.30 Exper. Ref: Hansch,C et al. (1995) Boiling Pt, Melting Pt, Vapor Pressure Estimations (MPBPWIN v1.42): Boiling Pt (deg C): 179.21 (Adapted Stein & Brown method) Melting Pt (deg C): -1.00 (Mean or Weighted MP) VP(mm Hg,25 deg C): 0.653 (Mean VP of Antoine & Grain methods) MP (exp database): -5 deg C BP (exp database): 188 deg C Water Solubility Estimate from Log Kow (WSKOW v1.41): Water Solubility at 25 deg C (mg/L): 8368 log Kow used: 1.30 (expkow database) no-melting pt equation used Water Sol Estimate from Fragments: Wat Sol (v1.01 est) = 19471 mg/L ECOSAR Class Program (ECOSAR v0.99h): Class(es) found: Aromatic Amines Henrys Law Constant (25 deg C) [HENRYWIN v3.10]: Bond Method : 2.22E-006 atm-m3/mole Group Method: 6.14E-006 atm-m3/mole Henrys LC [VP/WSol estimate using EPI values]: 1.141E-005 atm-m3/mole Log Octanol-Air Partition Coefficient (25 deg C) [KOAWIN v1.10]: Log Kow used: 1.30 (exp database) Log Kaw used: -4.042 (HenryWin est) Log Koa (KOAWIN v1.10 estimate): 5.342 Log Koa (experimental database): None Probability of Rapid Biodegradation (BIOWIN v4.10): Biowin1 (Linear Model) : -0.3491 Biowin2 (Non-Linear Model) : 0.0000 Expert Survey Biodegradation Results: Biowin3 (Ultimate Survey Model): 2.4117 (weeks-months) Biowin4 (Primary Survey Model) : 3.5925 (days-weeks ) MITI Biodegradation Probability: Biowin5 (MITI Linear Model) : 0.2741 Biowin6 (MITI Non-Linear Model): 0.0044 Anaerobic Biodegradation Probability: Biowin7 (Anaerobic Linear Model): 0.1765 Ready Biodegradability Prediction: NO Hydrocarbon Biodegradation (BioHCwin v1.01): Structure incompatible with current estimation method! Sorption to aerosols (25 Dec C)[AEROWIN v1.00]: Vapor pressure (liquid/subcooled): 79.3 Pa (0.595 mm Hg) Log Koa (Koawin est ): 5.342 Kp (particle/gas partition coef. (m3/ug)): Mackay model : 3.78E-008 Octanol/air (Koa) model: 5.4E-008 Fraction sorbed to airborne particulates (phi): Junge-Pankow model : 1.37E-006 Mackay model : 3.03E-006 Octanol/air (Koa) model: 4.32E-006 Atmospheric Oxidation (25 deg C) [AopWin v1.92]: Hydroxyl Radicals Reaction: OVERALL OH Rate Constant = 134.8403 E-12 cm3/molecule-sec Half-Life = 0.079 Days (12-hr day; 1.5E6 OH/cm3) Half-Life = 0.952 Hrs Ozone Reaction: No Ozone Reaction Estimation Fraction sorbed to airborne particulates (phi): 2.2E-006 (Junge,Mackay) Note: the sorbed fraction may be resistant to atmospheric oxidation Soil Adsorption Coefficient (PCKOCWIN v1.66): Koc : 72.53 Log Koc: 1.861 Aqueous Base/Acid-Catalyzed Hydrolysis (25 deg C) [HYDROWIN v1.67]: Rate constants can NOT be estimated for this structure! Bioaccumulation Estimates from Log Kow (BCFWIN v2.17): Log BCF from regression-based method = 0.301 (BCF = 2) log Kow used: 1.30 (expkow database) Volatilization from Water: Henry LC: 6.14E-006 atm-m3/mole (estimated by Group SAR Method) Half-Life from Model River: 101.6 hours (4.233 days) Half-Life from Model Lake : 1197 hours (49.86 days) Removal In Wastewater Treatment: Total removal: 2.27 percent Total biodegradation: 0.09 percent Total sludge adsorption: 1.83 percent Total to Air: 0.35 percent (using 10000 hr Bio P,A,S) Level III Fugacity Model: Mass Amount Half-Life Emissions (percent) (hr) (kg/hr) Air 0.179 1.9 1000 Water 45.6 900 1000 Soil 54.1 1.8e+003 1000 Sediment 0.105 8.1e+003 0 Persistence Time: 594 hr
Click to predict properties on the Chemicalize site
- 1-Click Docking
- 1-Click Scaffold Hop


